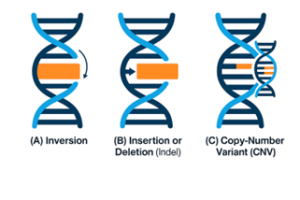
An increase in fillet yield is expected to translate into higher profit margins for fish farming enterprises selling mostly fillets. However, various technical issues make the genetic selection for the trait a challenge in breeding programmes. This may soon change with the adoption of genomic selection and artificial intelligence-based approaches to automate the quantification of body shape.
The fillet
Fish fillets comprise myotomal muscle fibres, connective tissue myosepta, blood vessels and various cell types involved in fat metabolism, immune function and growth. The meat mostly constitutes the white muscle used for rapid movements plus the relatively thin external layer of red muscle fibres used for slow swimming. Information on the development and growth of the fillet can be found here.
The proportion of fillet as a function of round or gutted weight (fillet yield) is not important when the species is sold as whole fish e.g., most breams and flatfish. However, for species primarily sold as fresh or frozen fillets increases in fillet yield are proportional to the revenue earned per metric tonne of production. In such cases, improving fillet yield leads to higher profit margins. This is particularly relevant for tilapia and catfish species where frozen fillets are globally traded commodities.
Fillet yield as a breeding trait
To achieve worthwhile genetic gain in any trait requires high-quality phenotyping. This involves precise definition of the trait and an ability to measure it consistently and accurately at scale and reasonable cost. Several types of fillet can be produced during secondary processing including the whole-cut, v-cut and j-cut (see here). In some cases, parts of the fillet (e.g. belly flaps) are trimmed (Fig. 1), the fillet can be skin-on or skin-off, and with Pangasius the layer of red muscle is largely removed.

Fig. 1. Trimmed fillet from a smoked Atlantic salmon (Salmo salar).
Larger fish inevitably have bigger fillets which will require additional feed, possibly more expensive diets and a longer production time. A selective breeding programme for this trait instead aims to increase the proportion of edible product relative to body weight without incurring higher production costs. Improvement in the proportion of fillet to body weight inevitably involves selection to reduce other non-edible anatomical features such as head size and/or volume of the visceral cavity.
Phenotyping
Fillet yield is most often expressed relative to head-on-gutted or whole (round) weight. When conducted by a skilled and trained individual, filleting can have the consistency and accuracy needed for genetic selection. The ratio of right and left fillet weight provides a good measure of consistency and can be used as a quality control filter. Since thousands of fillet yield measurements are required for most genetic applications, data collection requires careful planning and a dedicated team of filleters. The results of individual filleters and their performance over time should be taken into consideration in the final analysis of results.
For a trait to be improved by genetic selection it must be variable within the breeding population. Table 1 shows the published average and range of fillet yields for three species often sold as fillets. Average fillet yield varied from 36 to 57% with a range of 13-18% which is sufficiently variable for improvement by genetic selection.
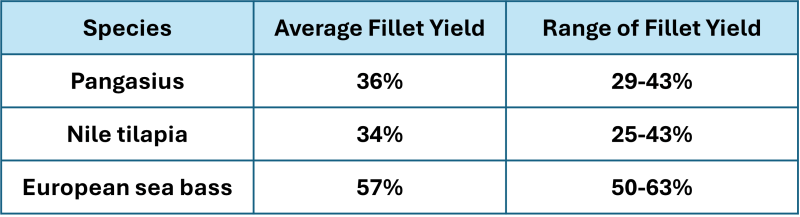
Table 1. Fillet yield expressed as a percentage of whole weight for three fish species.
In addition to being challenging to measure, relative fillet yield is a ratio with a high correlation between the numerator and denominator, and this presents difficulties for genetic analyses (see discussion below). Researchers have therefore explored correlated traits which are easier to measure accurately, are highly correlated with manual filleting and show moderate to high heritability. One of the most successful is headless gutted carcass weight (HGCW). For example, in rainbow trout HGCW shows a strong genetic correlation (0.97) with fillet yield, has high heritability (0.5), and can be more accurately measured in a commercial setting. Reduced operator error and the high heritability of HGCW was estimated to make indirect selection 50% more efficient than for fillet yield itself.
Heritability and potential for genetic improvement
Narrow sense heritability (h2) provides a measure of the potential for genetic improvement by selecting individuals based on their phenotypes. If h2 equals or exceed 0.2 then the trait is considered a good trait for improvement by selective breeding. It should be noted that h2 is population-specific and varies with developmental stage, environment and the design and management of the trials used for its measurement. Some studies have reported moderate to high values for the heritability of fillet yield including in rainbow trout (0.31), Nile tilapia (0.32), and common carp (0.5).
Ratio traits such as fillet yield may introduce biases which underestimate or overestimate genetic gain. Alternatives include the use of linear index theory or residual fillet weight (the residual of the regression of fillet weight on body weight, see here). Another possibility is to use waste weight (body weight – fillet weight) as the denominator rather than body weight because it is less correlated. Stochastic simulations using genetic parameters measured in sea bass, gilthead sea bream and rainbow trout populations revealed modest differences between the various methods with an average gain of +0.66% per generation using pedigree BLUP (Best Linear Unbiased Prediction) and classical selection.
Body shape measurements
Geomorphometric analysis can provide more sophisticated measures of body shape for indirect selection of fillet yield. Methods are based on the configuration of landmarks on the body surface obtained from 2D or 3D images. An example of the landmarks used for shape measurement in European sea bass is shown in Fig. 2.
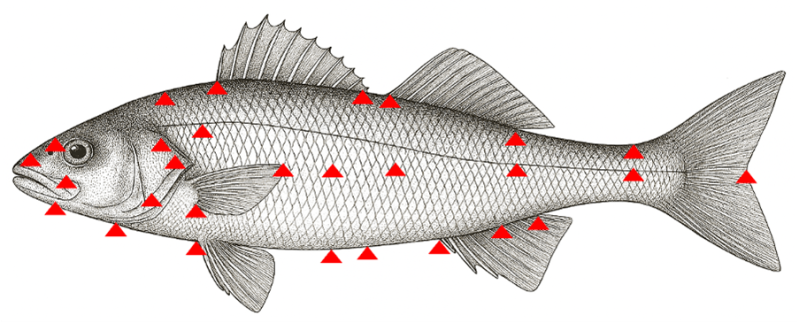
Fig. 2. Measuring size and shape by geomorphometric analysis. Landmarks for shape analysis in European sea bass shown in red. For further details see here.
The landmarks are scaled to unit centroid size and aligned by superimposition techniques designed to remove differences in position, orientation and size between individuals. Following further statistical analysis, the aligned shape co-ordinates can reliably discriminate between populations of the same species, but are still subject to inter-operator error (e.g., see here). The analysis pipeline for images requires skilled personnel, is time-consuming and therefore expensive. A recent study applied deep learning techniques to a data set of more than 2,000 wild and farmed gilthead seabream. A novel automated workflow for landmark placement achieved sub-millimetre accuracy reducing systematic and random digitisation errors relative to human annotations. Phenomics approaches such as these provide the scale and cost effectiveness needed for aquaculture breeding programmes and suggest the way forward. Moderate to high values (0.37-0.51) of narrow sense heritability were obtained for shape metrics at harvest size making this approach very promising for selective breeding (see here).
Internal morphology can be measured non-invasively using ultrasound. Fillet yield estimated using ultrasound has shown some potential for selection in rainbow trout but was less promising for Senegalese sole. In European sea bass a simple ratio of ultrasound signals for abdominal wall thickness to the depth of the peritoneal cavity had the highest genetic correlation with HGCW.
Design of the breeding programme
A major complication with classical selection is that fillet yield is a destructive trait which cannot be directly assessed in selection candidates. Fillet yield is therefore usually improved using sib selection or by selecting for correlated traits such as body shape metrics. Sib selection identifies the families with the highest fillet yields but ignores the considerable intra-family variation amongst the selection candidates.
Genomic selection (GS) which has become the standard for advanced breeding programmes is a game changer for improving fillet yield by selective breeding. It requires a training population of a few thousand individuals which encompasses the genetic variation of the selection candidates. The training population is phenotyped and genotyped at tens of thousands of genetic loci (usually with a 50-70K SNP chip) to develop a predictive model.
Genomic Estimated Breeding Values (GEBVs) are then obtained for selection candidates simply based on high density genotyping. For example, Garcia et al. selected rainbow trout using single step GBLUP (ssGBLUP) model for traits of fillet yield and body weight. ssGBLUP can include animals that only have phenotypes to increase the accuracy of the prediction model. In this case there were 12,890 phenotype records for fillet yield and body weight, with 2,484 fish genotyped with a 57K SNP chip. The resulting models increased prediction accuracy by up to 50% relative to pedigree BLUP. An associated Genome Wide Association Study (GWAS) found no significant Quantitative Trait Loci (QTLs). Previously, another GWAS on a different population of the same species identified two 20 SNP windows on chromosome 9 that explained 1.5% and 1.0% of the genetic variation in fillet yield.
These studies showed that fillet yield is a polygenic trait in rainbow trout for which GS models assuming equal SNP effects such as ssGBLUP are well suited. Leading up to the formation of Xelect, Tom Ashton co-founder of the company used a candidate gene approach involving multiple Atlantic salmon populations to discover and patent two SNPs associated with superior fillet yield. Individuals fixed for both favourable alleles had a field yield up to 2% higher than average. Smaller QTLs incorporated into weighted ssGBLUP models would be expected to make meaningful commercial gains in fillet yields over several generations. It is important that all associated genetic markers are validated in the target breeding population.
Future perspectives
One key question needs answering before deciding to incorporate any new trait into a breeding programme: “What is my likely return on investment (ROI)”?
In the case of fillet yield, recent developments in genomics have significantly increased the gain part of the equation. Several factors have also reduced the cost of achieving that gain. High density genotyping prices have reduced substantially over recent years. The ability to genotype more selection candidates for the same genotyping budget allows the breeding programme to operate at a higher selection pressure, further boosting the pace of genetic gain. It makes sense to add automated measurements of shape metrics when collecting genotype and phenotype information for the initial genomic selection training set. The adoption of a robust indirect selection index highly correlated with fillet yield in subsequent generations would serve to reduce costs further by reducing or even eliminating the amount of expensive manual filleting required.
All of these developments promise a more favourable ROI in the future. Now seems the perfect time to add fillet yield to the selection index, particularly for fish primarily sold as fillets, and species with short generation times such as Nile tilapia.